 
It is also available as part of several molecular graphics packages and websites. The source code of STRIDE has been freely accessible from the FTP server of the European Bioinformatics Institute since 1995 ( ). A weighted product of hydrogen bond energy and torsion angle probabilities for α-helix and β-sheet is used to determine the start and stop positions of secondary structure elements based on empirically optimized recognition thresholds. The hydrogen bond energy is calculated using an empirical energy function which takes into account the distance between the donor and the acceptor and the deviations from linearity of the bond angles ( 8, 9). The main difference between STRIDE and DSSP is that STRIDE considers both hydrogen bonding patterns and backbone geometry. It is thus a knowledge-based approach which uses, as training data, a carefully verified set of secondary structural elements defined by crystallographers who have deposited structures in the Protein Data Bank (PDB) ( 7). 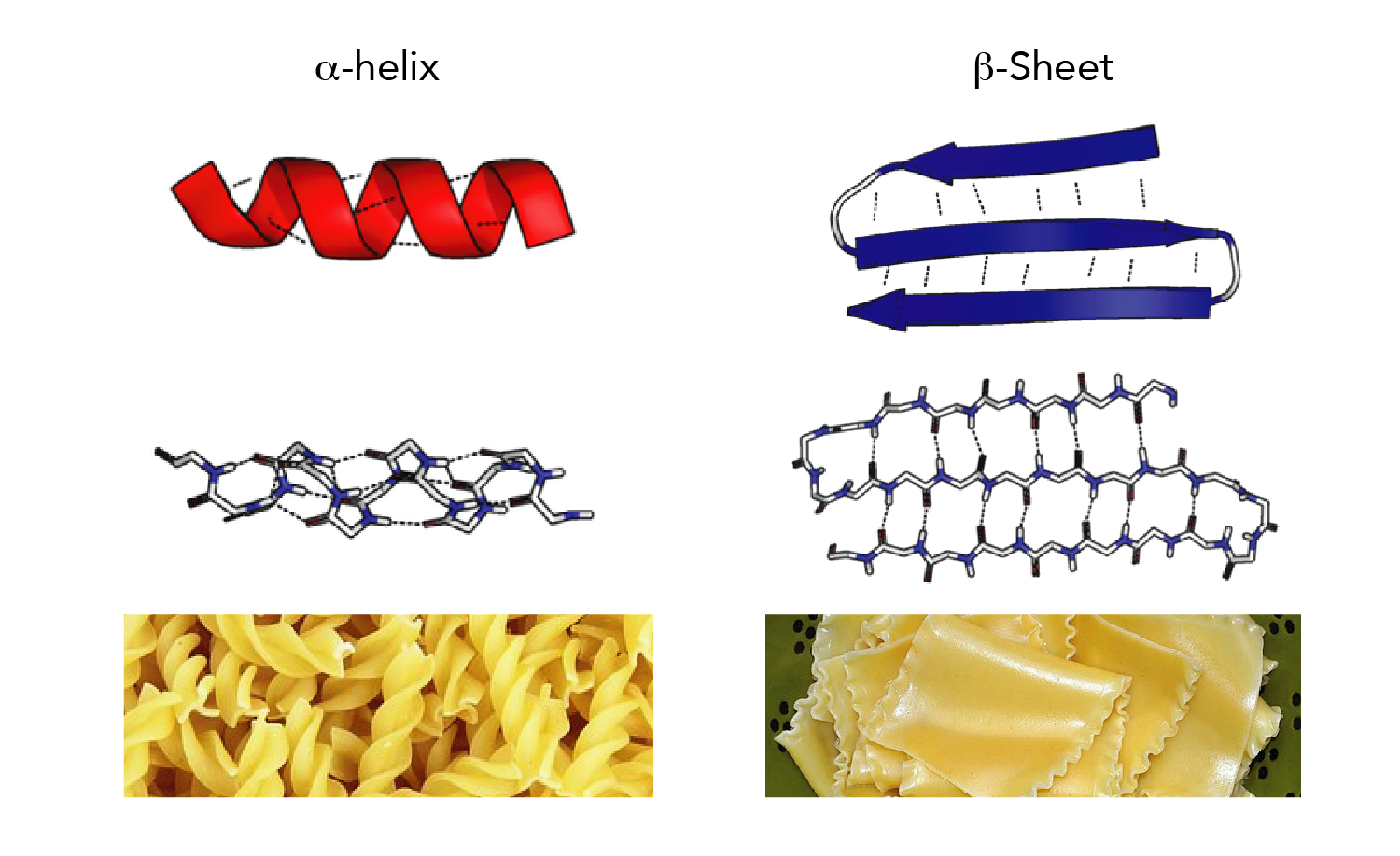
Our method, STRIDE ( 6), was developed with a specific goal to accurately reproduce secondary structure designations created by human experts. More recently, an improved version of DSSP, called DSSPcont, has been developed which takes into account the structural variations in proteins ( 5). The DEFINE algorithm ( 3) compares inter-atomic distance matrices of structural fragments to idealized reference distance masks typical for a particular secondary structure type, while P-Curve ( 4) is based on quantification of backbone curvature using differential geometry. Hydrogen bonds between peptide units are assigned if the electrostatic interaction energy between C=O of one residue and N–H of another residue is <−0.5 kcal/mole. The most widely used method, DSSP ( 2), defines secondary structure elements as repeating elementary hydrogen bonded patterns. Several automatic tools for secondary structure assignment from known atomic coordinates are available. To a large extent, our visual notion of proteins is based on cartoon diagrams showing α-helices and β-strands as cylinders and arrows, respectively. 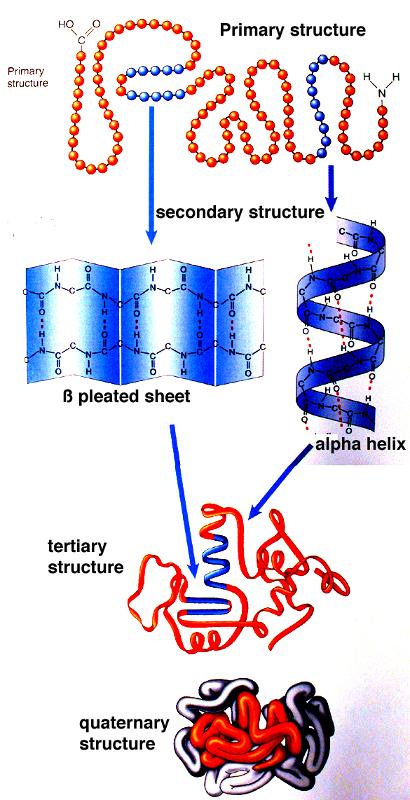
It serves as a basis for virtually all subsequent analyses, including visualization, structure comparison and classification, homology modelling, threading and sequence alignment. Identification of secondary structure elements is a major step in the characterization of a newly determined protein structure. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
